Research
Research Themes
My research primarily focuses on exploring complex systems and understanding how information processing occurs in chemical and physical systems. The aim is to understand the emergence of living systems from non-living matter, discover novel functional materials, and develop novel computational frameworks. This requires integrating various disciplines including physics, chemistry, computer science, and engineering. The four key topics of my research are Physics of life and matter, Unconventional Computation, Automated Materials discovery using self-driving labs, and Microsystems Chemistry and Microtechnology.
Physics of Matter and Life
The current physics does not have a notion of function and emergence of novelty which are the key principles governing biological laws. We develop Assembly Theory (AT) as a potential approach to unify Physics and Biology to explain evolution and complexity. My research focuses on developing the fundamentals of Assembly Theory to describe selection and combine it with laws of Thermodynamics and Statistical Mechanics. Additionally, I also design experiments to observe and quantify selection using AT in complex chemical systems and utilise various spectroscopic techniques to detect molecular biosignatures as life-detection. The key questions can be defined as follows,
- How does life originate from non-living matter?
- How fixed and timeless laws of physics lead to the dynamic laws of biological systems?
- How does selection emerge in a non-biological system and how can it be quantified?
- How to detect life elsewhere in the universe using molecules as biosignatures?
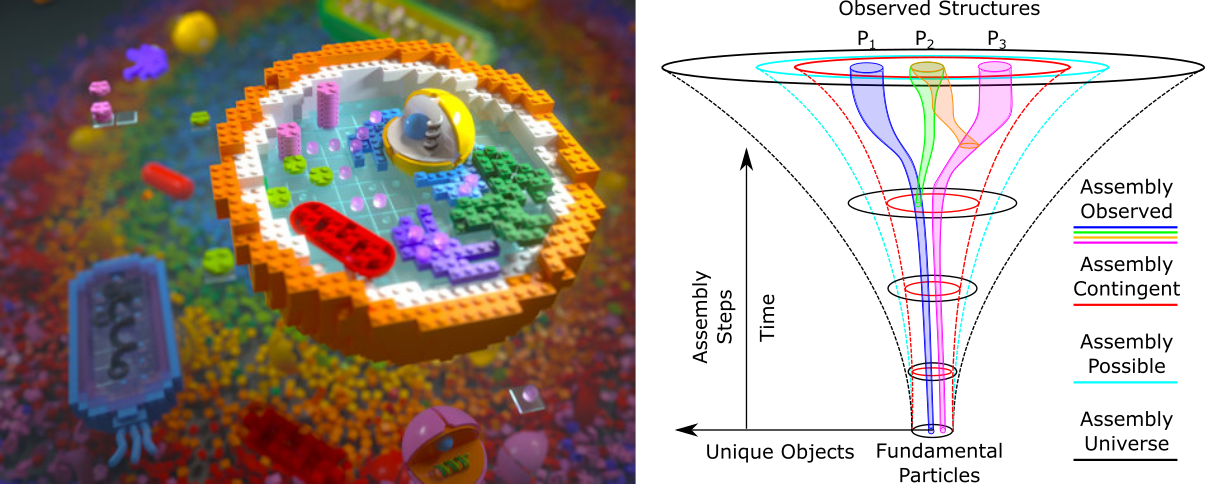
Fig 1. Left: Artistic representation of a cell showing self-assembled individual components within a vast combinatorial universe. (Image Credit: Dr Anna Tanczos, Sci-Comm Studios) Right: Nested structure of combinatorial assembly spaces, where different Assembly Contingent and Assembly Possible represent directed and undirected dynamics in forward processes respectively. (Nature 2023, 622 (7982), 321–328)
The Assembly Theory was recently published in Nature, and our novel strategy to detect life using biosignature has been published in ACS Central Science. See publication list complete list of publications. Here are the key publications,
(1) Sharma, A.; Czégel, D.; Lachmann, M.; Kempes, C. P.; Walker, S. I.; Cronin, L. Assembly Theory Explains and Quantifies Selection and Evolution. Nature 2023, 622 (7982), 321–328. PAPER
(2) Jirasek, M.; Sharma, A.; Bame, J. R.; Mehr, S. H. M.; Bell, N.; Marshall, S. M.; Mathis, C.; MacLeod, A.; Cooper, G. J. T.; Swart, M.; Mollfulleda, R.; Cronin, L. Investigating and Quantifying Molecular Complexity Using Assembly Theory and Spectroscopy. ACS Cent. Sci. 2024, 10 (5), 1054–1064. (shared first author) PAPER
Some recent reviews on theoretical and experimental aspects of Assembly Theory.
(1) Ellis, G. F. R. How Purposeless Physics Underlies Purposeful Life. Nature 2023, 622 (7982), 247–249. PAPER
(2) Slocombe, L.; Walker, S. I. Measuring Molecular Complexity. ACS Cent. Sci. 2024, 10 (5), 949–952. PAPER
Unconventional Computation
The exponential growth of computing power is getting restricted due to fabrication limitations at the nanoscale as well as von Neumann bottleneck which limits the data throughput between processor and memory. Additionally, the scaling of deep learning models is constrained by the energy efficiency of current computational architectures. My research focuses on developing hybrid computational architectures utilizing “the best of both worlds” approach by combining electronic digital logic with probabilistic logic from molecular and physical systems. Recently, we have utilized the Belousov-Zhabotinsky (BZ) reaction and combined with digital logic, developed a novel computing architecture capable of solving combinatorial optimization problems efficiently. We also developed physically instantiated Cellular Automata inspired by Conway’s Game of Life. To demonstrate the scalability, we also miniaturized the architecture using electrode arrays with pH-responsive thin gel films. The key questions my research focuses on are,
- How to program physical and chemical systems spatiotemporally to develop efficient computational architecture?
- How to explore and employ non-linear dynamics in physical systems to develop computers capable of performing complex machine learning tasks energy efficient?
- Designing hybrid logic for digital-analog distributed computation.
Our BZ computer was recently published in Nature Communications and our scalable classical-molecular computer was published in ACS Central Science. See the publication list for complete list of publications.
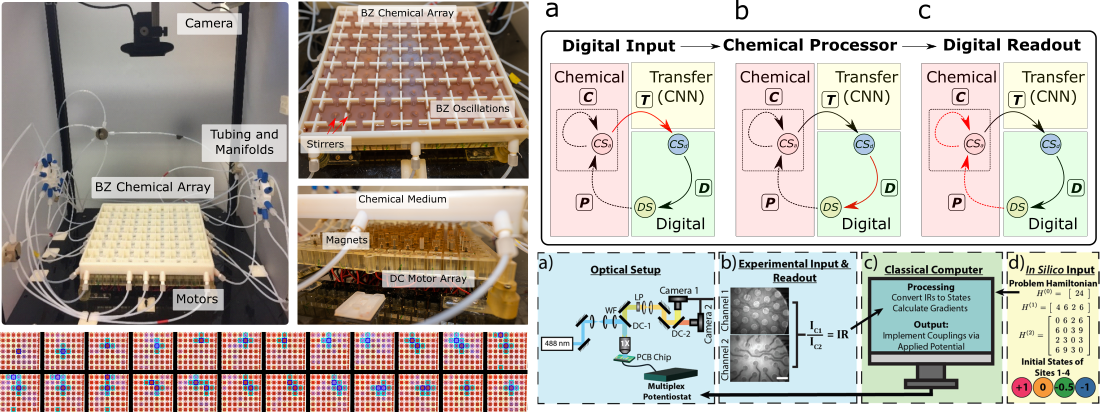
Fig 2. Left: BZ Computer experimental setup and demonstrating “BZ Game of Life” Cellular Automata showing emergent life-like behaviour as an example of hybrid computation. (Nat Comm 2024, 15 (1), 1984) Right: Top image shows hybrid computer informational loop where spatiotemporal spatiotemporal states dynamically propagates between digital and chemical domains (Nat Comm 2024, 15 (1), 1984). Bottom image shows implementation of hybrid classical molecular computer using pH sensitive gel over a fully addressable electrode array. (ACS Cent. Sci. 2023, 9 (7), 1453–1465)
Key Publications
(1) Sharma, A.; Ng, M. T.-K.; Parrilla Gutierrez, J. M.; Jiang, Y.; Cronin, L. A Programmable Hybrid Digital Chemical Information Processor Based on the Belousov-Zhabotinsky Reaction. Nat Commun 2024, 15 (1), 1984. PAPER
(2) Krasecki, V. K.; Sharma, A.; Cavell, A. C.; Forman, C.; Guo, S. Y.; Jensen, E. T.; Smith, M. A.; Czerwinski, R.; Friederich, P.; Hickman, R. J.; Gianneschi, N.; Aspuru-Guzik, A.; Cronin, L.; Goldsmith, R. H. The Role of Experimental Noise in a Hybrid Classical-Molecular Computer to Solve Combinatorial Optimization Problems. ACS Cent. Sci. 2023, 9 (7), 1453–1465. PAPER
Automated Materials Discovery using Self-driving Labs
The discovery of novel materials involves exploration of high-dimensional chemical space which includes chemical reagents, process variables, and product analysis which is a laborious task. My research focuses on developing robotic platforms using a DIY approach for discovering novel inorganic materials and multi-component systems by utilizing a closed-loop approach with state-of-the-art machine learning algorithms. Recently, we have developed an autonomous closed-loop nanomaterials discovery robot with programmable synthetic language (XDL) and by combining experimental and theoretical frameworks, we demonstrate the exploration and optimization of highly reproducible novel metallic nanomaterials. The discovered experimental protocols are digitizable and can be performed reproducibly independent of any specific experimental system. We also develop closed-loop formulation robots for protocells (self-propelling droplets) research towards artificial life. The robot explores the vast combinatorial space of oil formulations searching for novelty in the dynamics of droplets and optimization for life-like properties. The key questions on which my research focuses on are,
- Efficient exploration and optimization of vast chemical space by combining robotic platforms and spectroscopic techniques to discover novel inorganic and nanomaterials.
- Digitization of synthetic procedures for reproducibility.
- Exploring principles of life in multiphase self-propelling droplets using robotic assistant.
Our nanomaterials discovery and optimization robot as well as formulation robot both were recently published in Science Advances. We also developed a fast GPU-accelerated algorithm to simulate the spectral properties of dynamically evolving nanomaterials. See publications for the complete list.
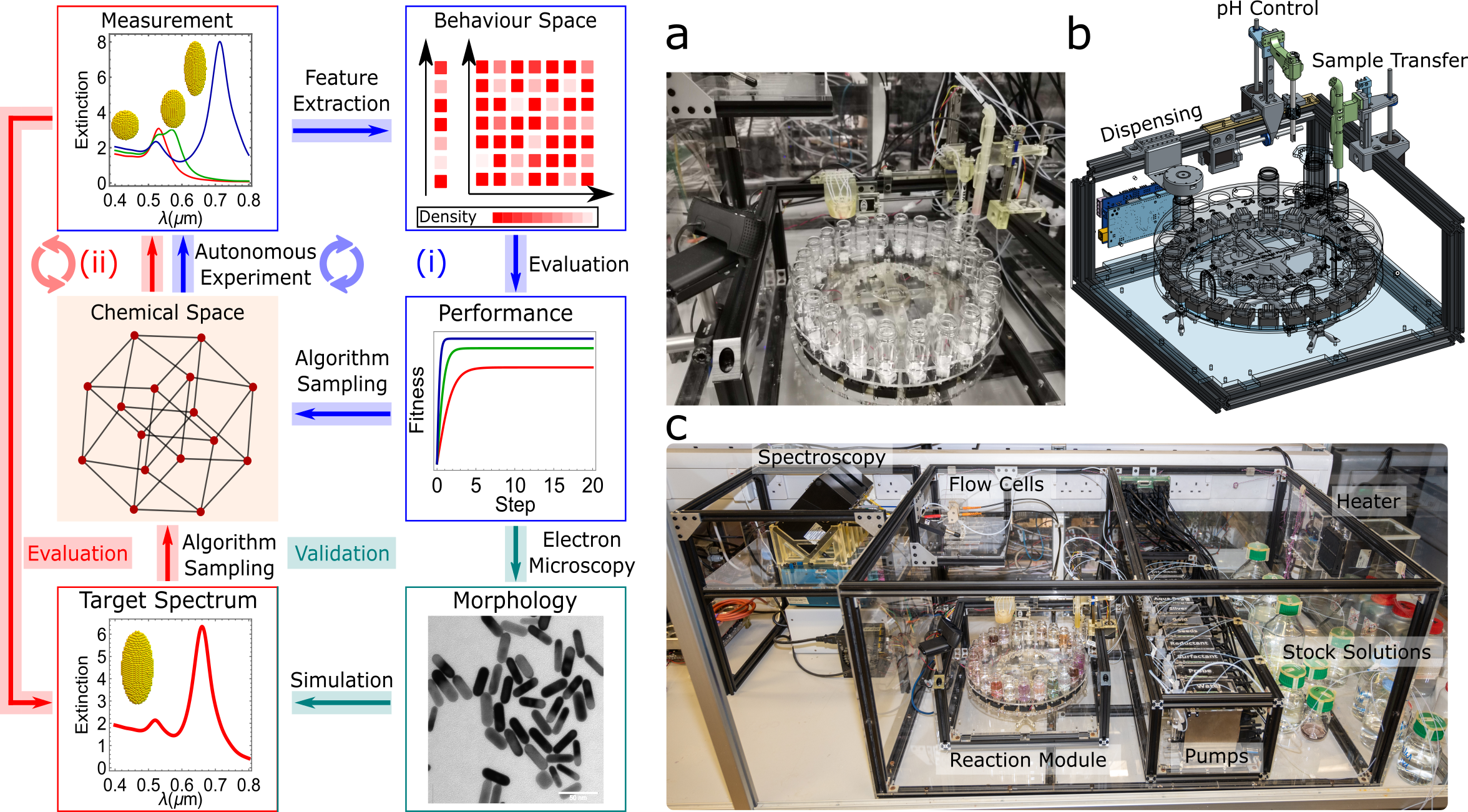
Fig 3. Left: Experimental concept for closed-loop nanomaterials discovery robot. Right: Experimental implementation of nanomaterials discovery robot comprising chemical reaction module, pumps control, flow cells and spectroscopy for closed-loop approach (Sci. Adv. 2022, 8 (40), eabo2626).
Key Publications
(1) Jiang, Y.; Salley, D.; Sharma, A.; Keenan, G.; Mullin, M.; Cronin, L. An Artificial Intelligence Enabled Chemical Synthesis Robot for Exploration and Optimization of Nanomaterials. Sci. Adv. 2022, 8 (40), eabo2626. PAPER
(2) Jiang, Y.; Sharma, A.; Cronin, L. An Accelerated Method for Investigating Spectral Properties of Dynamically Evolving Nanostructures. J. Phys. Chem. Lett. 2023, 14 (16), 3929–3938. (shared first author) PAPER
(3) Grizou, J.; Points, L. J.; Sharma, A.; Cronin, L. A Curious Formulation Robot Enables the Discovery of a Novel Protocell Behavior. Sci. Adv. 2020, 6 (5), eaay4237. PAPER
Microsystems Chemistry and Microtechnology
Programmable nanoscale Lab-on-a-Chip devices are very useful for cost-efficient and massively parallel biomolecular processing at low volume and low concentration regimes. My research focuses on designing PDMS-based microfluidic devices mapped to a high-density microelectrode array to perform on-chip generation of DNA libraries in nanolitre droplets together with electrophoresis and isotachophoresis. I also used theoretic and computational tools to understand diffuse charge dynamics in microelectrochemical systems. We also developed autonomous microscale “particles” equipped with programmable CMOS electronics (called as lablets), energy source, sensors, actuators, and digital logic to perform localized information processing in chemical medium towards developing artificial cells, distributed computational frameworks etc. The key questions on which my research focuses on are,
- Programmable massively parallel exploration of chemical reaction spaces in nanolitre droplets with microfluidic control.
- Diffuse charge dynamics and programmability of non-linear electrochemical dynamics at microscale.
- Information processing and bio-inspired swarm dynamics in autonomous microscale devices towards developing artificial cells.
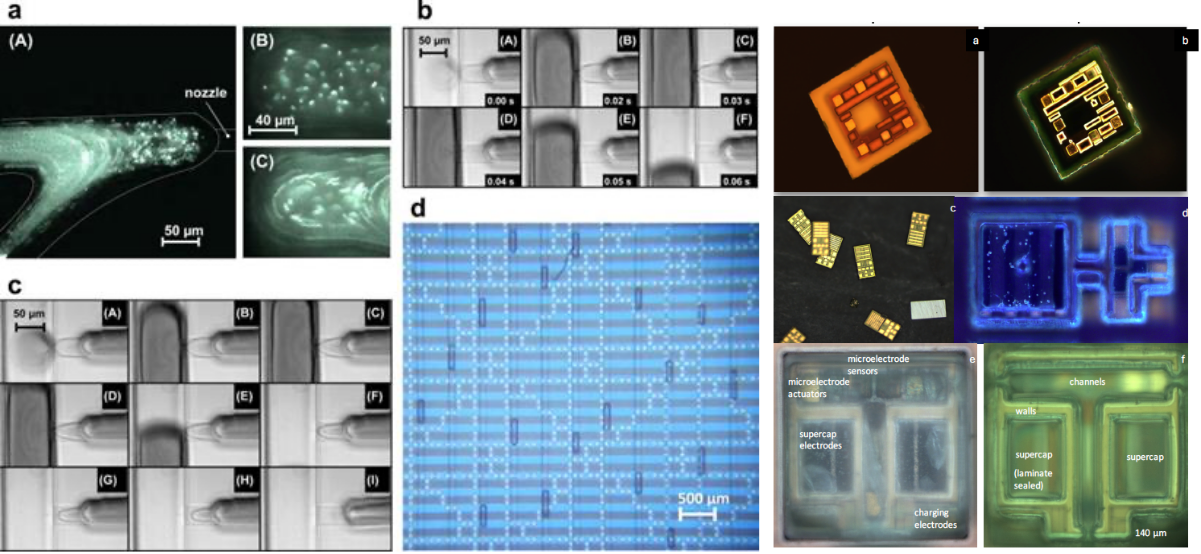

Fig 4. Left: A-D shows nanolitre on-demand droplet generation and mixing in a microfluidic device. The architecture is highly programmable and optimizable based on experimental requirements and has been used in generating DNA libraries at nanoscale (Biomicrofluidics 2015, 9 (1), 014119, Biomicrofluidics 9, 044103 2015). Right: A-F shows images of autonomous microscale CMOS electronic devices with on-chip supercapacitors, microchannels for electroosmotic drive, and sensors.
Key Publications
(1) Tangen, U.; Sharma, A.; Wagler, P.; McCaskill, J. S. On Demand Nanoliter-Scale Microfluidic Droplet Generation, Injection, and Mixing Using a Passive Microfluidic Device. Biomicrofluidics 2015, 9 (1), 014119. PAPER
(2) Sharma, A.; McCaskill, J. S. Autonomous Lablet Locomotion and Active Docking by Sensomotory Electroosmotic Drive. In 07/20/2015-07/24/2015 In Proceedings of the Artificial Life Conference 2016; The MIT Press, 2015; pp 456–462. PAPER
(3) Maeke, T.; McCaskill, J.; Funke, D.; Mayr, P.; Sharma, A.; Tangen, U.; Oehm, J. Autonomous Programmable Microscopic Electronic Lablets Optimized with Digital Control. arXiv 2024. PAPER
